Note
Click here to download the full example code
Plot confusion matrix¶
Plot confusion matrix from Cross-Validation, with F1 as subplot.
Import librairies¶
from museotoolbox.ai import SuperLearner
from museotoolbox.cross_validation import RandomStratifiedKFold
from museotoolbox.charts import PlotConfusionMatrix
from museotoolbox import datasets
from sklearn.ensemble import RandomForestClassifier
Load HistoricalMap dataset¶
X,y = datasets.load_historical_data(low_res=True,return_X_y=True)
Create CV¶
RSKF = RandomStratifiedKFold(n_splits=2,
random_state=12,verbose=False)
Initialize Random-Forest¶
classifier = RandomForestClassifier()
Start learning¶
SL = SuperLearner(classifier=classifier,param_grid=dict(n_estimators=[10,50]))
SL.fit(X,y,cv=RSKF)
Get kappa from each fold¶
for stats in SL.get_stats_from_cv(confusion_matrix=False,kappa=True):
print(stats['kappa'])
Out:
0.925091417957069
0.9164468361024596
Get each confusion matrix from folds¶
cms = []
for stats in SL.get_stats_from_cv(confusion_matrix=True):
cms.append(stats['confusion_matrix'])
print(stats['confusion_matrix'])
Out:
[[934 12 0 1 0]
[ 29 258 0 2 0]
[ 1 0 283 0 0]
[ 1 13 3 49 0]
[ 0 1 0 0 0]]
[[933 13 0 1 0]
[ 28 247 0 14 0]
[ 0 0 284 0 0]
[ 0 10 1 55 0]
[ 1 0 0 0 0]]
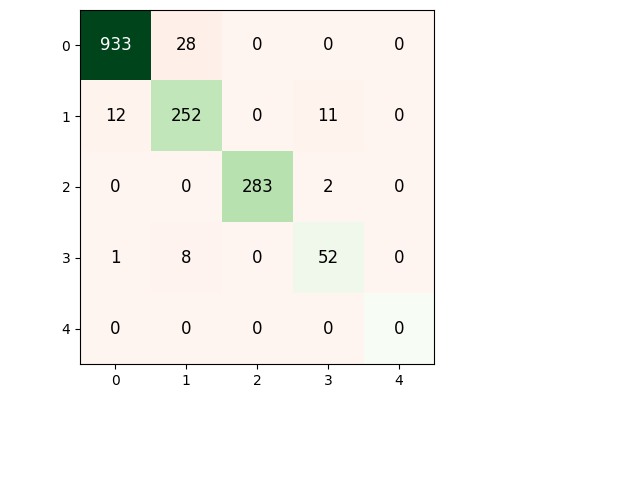
Plot confusion matrix¶
import numpy as np
meanCM = np.mean(cms,axis=0).astype(np.int16)
pltCM = PlotConfusionMatrix(meanCM.T) # Translate for Y = prediction and X = truth
pltCM.add_text()
pltCM.color_diagonal()
Total running time of the script: ( 0 minutes 1.013 seconds)